A general conversion program is available at READSEQ. Most web-based tools allow a number of input and output formats, such as FASTA format and GenBank format however, the use of specific tools authored by individual research laboratories can be complicated by limited file format compatibility. Sequence alignments can be stored in a wide variety of text-based file formats, many of which were originally developed in conjunction with a specific alignment program or implementation. For multiple sequences the last row in each column is often the consensus sequence determined by the alignment. In protein alignments, such as the one in the image above, colour is often used to indicate amino acid properties to aid in judging the conservativeness of a given amino acid substitution. Many sequence visualization programs also use color to display information about the properties of the individual sequence elements in DNA and RNA sequences, this equates to assigning each nucleotide its own color. As in the image above, an asterisk or pipe symbol is used to show identity between two columns other less common symbols include a colon for conservative substitutions and a period for semiconservative substitutions. In text formats, aligned columns containing identical or similar characters are indicated with a system of conservation symbols. In almost all sequence alignment representations, sequences are written in rows arranged so that aligned residues appear in successive columns. A variety of computational algorithms have been applied to the sequence alignment problem, including slow but formally optimizing methods like dynamic programming and efficient heuristic or probabilistic methods designed for large-scale database search.Īlignments are commonly represented both graphically and in text format. Local alignments are often preferable, but can be more difficult to calculate because of the additional challenge of identifying the regions of similarity. By contrast, local alignments identify regions of similarity within long sequences that are often widely divergent overall. 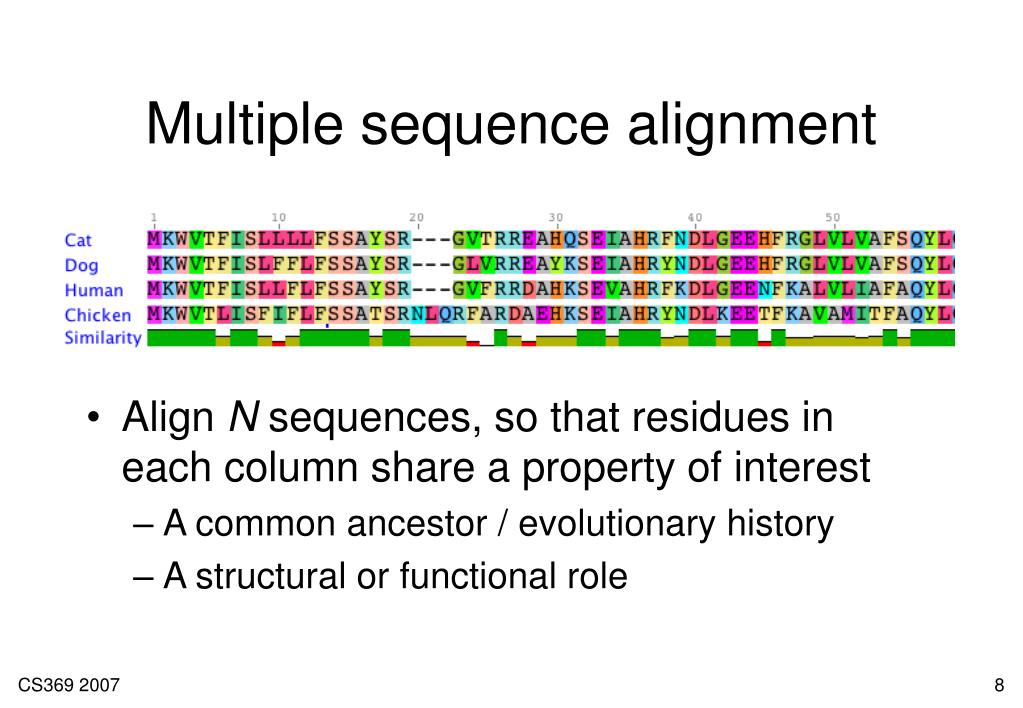
Calculating a global alignment is a form of global optimization that "forces" the alignment to span the entire length of all query sequences. Computational approaches to sequence alignment generally fall into two categories: global alignments and local alignments. Instead, human knowledge is primarily applied in constructing algorithms to produce high-quality sequence alignments, and occasionally in adjusting the final results to reflect patterns that are difficult to represent algorithmically (especially in the case of nucleotide sequences). Very short or very similar sequences can be aligned by hand however, most interesting problems require the alignment of lengthy, highly variable or extremely numerous sequences that cannot be aligned solely by human effort. In business, more specifically in marketing, sequences of purchases are also increasingly being analyzed by the same methods as in bioinformatics. Sequence alignment can be used for non-biological sequences, such as identifying similarities in a series of letters and words present in human language. Although DNA and RNA nucleotide bases are more similar to each other than to amino acids, the conservation of base pairing can indicate a similar functional or structural role. The absence of substitutions, or the presence of only very conservative substitutions (that is, the substitution of amino acids whose side chains have similar biochemical properties) in a particular region of the sequence, suggest that this region has structural or functional importance. In protein sequence alignment, the degree of similarity between amino acids occupying a particular position in the sequence can be interpreted as a rough measure of how conserved a particular region or sequence motif is among lineages. If two sequences in an alignment share a common ancestor, mismatches can be interpreted as point mutations and gaps as indels (that is, insertion or deletion mutations) introduced in one or both lineages in the time since they diverged from one another. A sequence alignment, produced by ClustalW between two human zinc finger proteins identified by GenBank accession number. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
